I would like to use this thread to make announcements regarding OHDSI Phenotype Library. The announcements will be on three topics
- Atlas-Phenotype is maintained by the OHDSI Phenotype Development and Evaluation Workgroup. This workgroup will use this thread to make any key announcements regarding cohort definitions.
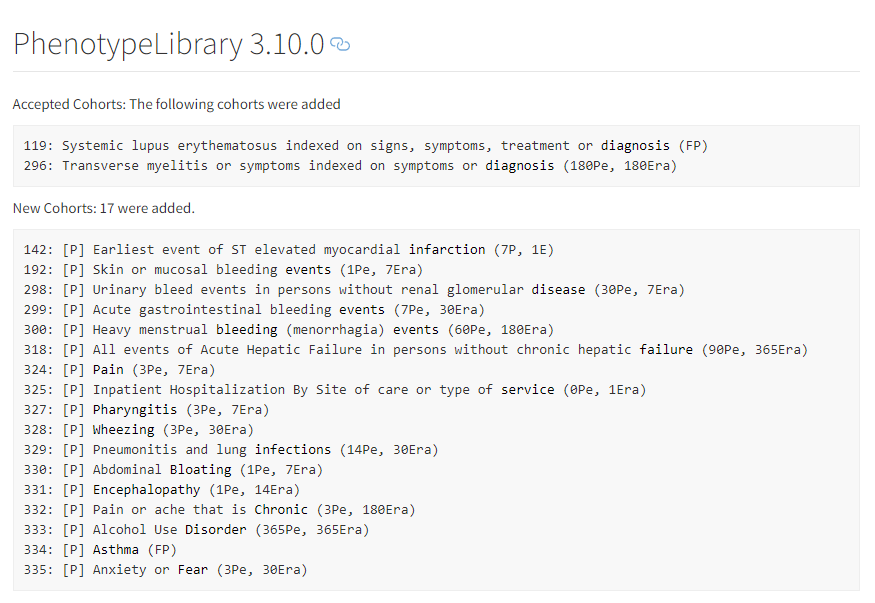
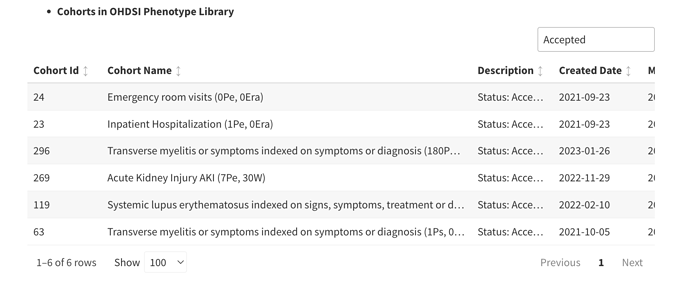
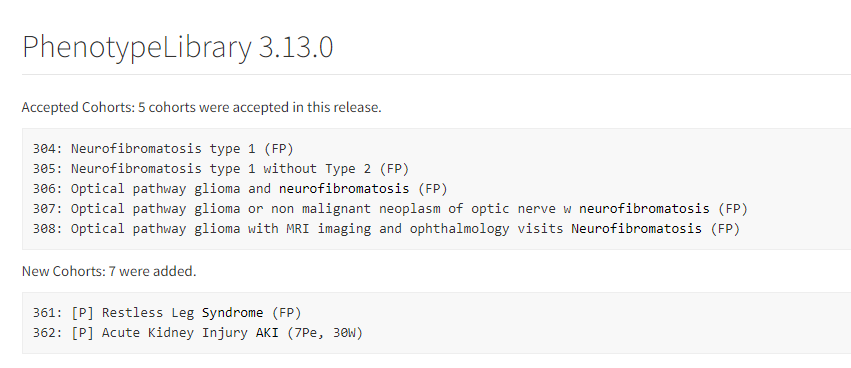
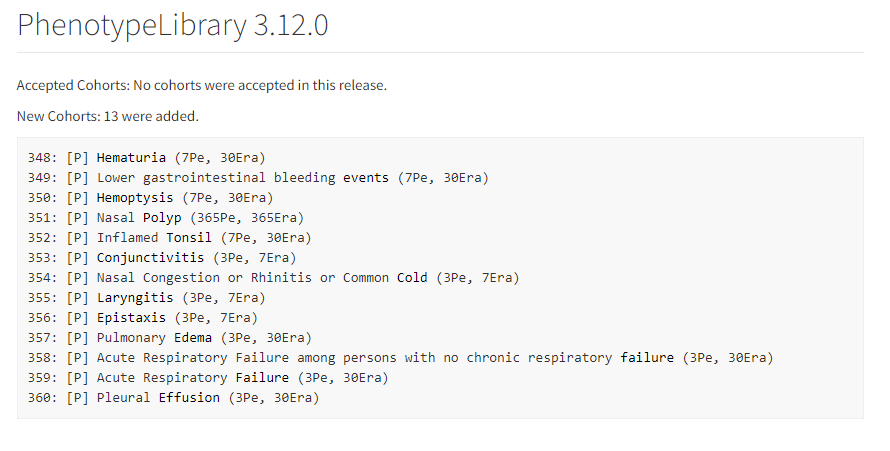
- OHDSI/PhenotypeLibrary : Periodically, cohort definitions from Atlas-Phenotype will be published into this repository. This will be version controlled with HADES compatibility. It will be an installable, referenceable, non executable R package with HADES version numbering convention.
- ohdsi-studies/PhenotypeLibraryDiagnostics: An executable study package called PhenotypeLibraryDiagnostics will be maintained, and it will be dependent on OHDSI/PhenotypeLibrary . Data partners will be pasked to execute this package on their sites and contribute the results back to the community. The results will be compiled and uploaded to data.ohdsi.org/PhenotypeLibrary (thank you @admin for support on this)
Submissions to the OHDSI Phenotype Library
- The OHDSI Phenotype Development and Evaluation Workgroup is considering submissions to the OHDSI Phenotype Library.
- Valid cohort definition submissions that originate from an Atlas (or Circe compliant source like CapR will go into [submitted] state and will be available at Atlas-Phenotype. These are not considered [accepted]. To be accepted, they should complete an evaluation by the Phenotype Development and Evaluation Workgroup and only accepted Cohort Definitions get promoted to the version controlled Library repository OHDSI/PhenotypeLibrary. Cohort Definitions that do not conform to Circe (e.g. probabilistic definitions and SQL only definitions) are also accepted.
- We encourage you to submit the results of running CohortDiagnostics on the submitted OHDSI CohortDefinitionSet . Other data partners will be invited to run CohortDiagnostics on the submitted cohortDefinitionSet. Based on the collective review of the CohortDiagnostics output a decision of acceptance will be made.
Technical Requirements:
- Submissions should conform to the OHDSI CohortDefinitionSet object as defined by OHDSI/CohortGenerator. (example: use ROhdsiWebApi::exportCohortDefinitionSet ) . This is required to make definitions importable into Atlas-Phenotype .