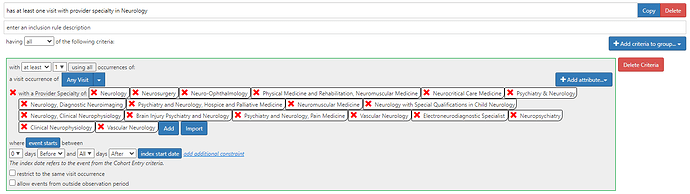
I’d like to introduce thoughts from our joint team (at UCLA and Northwestern University) on Parkinson’s disease and parkinsonism. Thanks to @Patrick_Ryan for encouragement to participate even as we have yet to setup our own OMOP-CDM instance.
Goal: to develop and evaluate algorithms (and impact of criteria used within them) that assess incidence and prevalence of
- Parkinson’s disease (PD) and
- neurodegenerative parkinsonism disorders (PD included)
Clinical description
Parkinson’s disease (PD) is a neurodegenerative disease that has been estimated to be increasing in prevalence based on large scale epidemiologic work. It is the most frequent form of neurodegenerative parkinsonism, itself a subset of parkinsonism syndromes. More details below.
PD is considered the condition when there is specific degeneration of the substantia nigra dopaminergic (DA) neurons over time producing the disease. It is estimated that well over 50% of DA cells will degenerate before clinical symptoms appear. This is the classic and core definition; however, increasingly there is appreciation of that PD itself is a multisystem disorder with involvement of other neurotransmitters and other systems within and not within the nervous system (often noted as non-motor symptoms – constipation, cognitive impairment, sleep disorders, autonomic dysfunction, depression, anxiety etc).
PD is the most common form of neurodegenerative parkinsonism (80-85%).
Neurodegenerative parkinsonisms (other than PD) are defined by having parkinsonism plus other neurologic systems that are affected by degeneration besides the specific PD degeneration described above. There will be described in more detail below – and include progressive supranuclear palsy (PSP), multiple systems atrophy (MSA), corticobasal degeneration (CBD) and others. Frequently, patients with neurodegenerative parkinsonism may be diagnosed with PD early on before neurodegenerative features become sufficiently prominent for clinical diagnosis over time. If patients have a clear diagnosis of PSP or MSA (for example), those are not considered PD, but are within neurodegenerative parkinsonism.
A good reference and overview of PD as a clinical syndrome, its early diagnosis, and its evolution over time is Armstrong and Okun (2020 Diagnosis and Treatment of Parkinson Disease: A Review - PubMed )
Clinical diagnosis of Parkinson’s disease (PD)
Parkinson’s disease is a clinical diagnosis during life with no lab/imaging studies that definitively establish the diagnosis. Parkinson’s disease is the most common form of parkinsonism.
Parkinsonism is defined as the syndrome of
- rigidity
- rest tremor
- bradykinesia (slowness of movement)
- postural instability
Each of above can be recorded in a variety of different ways – and (except for rest tremor) tend to be highly nonspecific. The well-established practical UK Brain Bank criteria for parkinsonism is that 2 of the above 4 findings are present with at least 1 of the two being either rigidity or bradykinesia. Note, tremor is not required. The current Movement Disorders 2015 consensus criteria defines parkinsonism as bradykinesia with or without rest tremor, rigidity, or both – and of course – must not be attributable to other causes (e.g. slowness from depression, drugs, illness, catatonia, lack of motivation, other neurologic disorders). Many of these criteria are often considered routinely impractical outside a movement disorders specialty evaluation/setting.
Parkinson’s disease is clinically diagnosed when patients have parkinsonism and
- has supportive criteria without significant red flags or confounding findings
Supportive criteria generally are accepted as - clear and dramatic beneficial response to dopaminergic medication – see below
- presence of levodopa-induced dyskinesia
- gradual progression over time
Red flags and confounding findings include - early (typically within 3 years of onset of sx) severe orthostatic hypotension
- early (within 3 years…) severe urinary incontinence/retention, not otherwise explained
- early recurrent falls (within first 3 years of sx)
- early severe dementia (controversial) – some criteria consider PD occurring independent of dementia; others will exclude early dementia or certain types of dementia
- complete absence of parkinsonism progression after 5 years
- neurologic findings such as ataxia, spasticity, aphasia, apraxia, supranuclear gaze palsy
- normal DATscan (dopamine transporter scan)
- absence of secondary causes for parkinsonism:
-
- drug-induced parkinsonism (typically has exposure to medication at the time parkinsonism is being diagnosed or considered and improves or does not progress if med is reduced, discontinued)
-
- multi-infarct vascular parkinsonism
-
- normal pressure hydrocephalus
-
- recurrent head trauma, severe traumatic brain injury or concussion with parkinsonism
-
- history of encephalitis with subsequent parkinsonism
The beneficial response to dopaminergic medication is considered highly specific, but a failure of response is difficult to prove as most criteria recommend 1000 mg/day levodopa be used without response before definitive criteria of not responding is registered. It is noted that this criteria makes having the prescription of a PD med not specific to making the diagnosis because it is not infrequently used as a “test” prescription to see if the patient will respond or not.
The progression of symptoms is slow and it may take several years before a clinically probable PD case is diagnosed. Quality measures for assessing quality care of neurologists recommend annual re-review of diagnosis for the first 5 years after symptoms.
Conversely, many of the features that define neurodegenerative (non-PD) early in the course of disease (to be described below) become common in late-stage typical PD and one must be cautious not to overdiagnose neurodegenerative (non-PD) parkinsonism if it is clear that one has typical PD but that the patient has simply developed the other neurologic features as part of advanced late-stage PD disease.
Clinical diagnosis of neurodegenerative parkinsonisms (PD is one of them)
Non-neurodegenerative parkinsonism (secondary parkinsonism):
These should be excluded from cohorts of PD and neurodegenerative parkinsonism.
To distinguish neurodegenerative parkinsonism (of interest) from non-neurodegenerative parkinsonism can be challenging, especially since two categories may co-exist or be considered/coded early and over time, clinical clarity will hopefully emerge.
Typically, if a clinical diagnosis of a clearly established non-neurodegenerative parkinsonism is made and that diagnosis correlates with the clinical course over time for that particular etiology (varies by etiology) and which fully explains the neurologic presentation, then the likelihood of a neurodegenerative parkinsonism is considered less likely.
Having a non-neurodegenerative parkinsonism (NNP) of course does not exclude PD (or other neurodegenerative parkinsonism) as the etiology may have unmasked an incipient PD which would be detected by abnormal clinical course that deviates from the NNP etiology over time.
These non-neurodegnerative parkinsonisms (by etiology) are often referred “secondary parkinsonisms” as some degree of warning/exclusionary criteria. Common ones are vascular (multistroke) parkinsonism and drug-induced parkinsonism. Others described below.
Neurodegenerative parkinsonism (non-PD):
Among neurodegenerative parkinsonism, above clinical guidelines help distinguish PD from non-PD neurodegenerative parkinsonism.
The non-PD neurodegenerative parkinsonisms are defined by degeneration of other neurologic systems besides the specific PD pathology. Neurodegenerative parkinsonisms tend to progress over time, so time again becomes a distinguishing element as features below emerge gradually and can be nonspecific or not routinely assessed well enough to be difficult to recognize are present in an EHR.
Common neurodegenerative parkinsonisms (besides PD) and their associated symptoms / neurologic system involved:
-
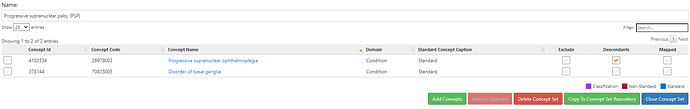
PSP – progressive supranuclear palsy – parkinsonism + early falls & supranuclear gaze palsy
-
CBD – corticobasal degeneration – park-ism + asymmetric dystonia, apraxia, alien-limb
-
MSA – multiple systems atrophy – park-ism + autonomic dysfunction (bladder/orthostasis)
-
MSA-c – multiple systems atrophy – MSA + cerebellar ataxia
-
MSA-p – multiple systems atrophy – MSA + parkinsonism predominant subtype
-
SDS – Shy Drager Syndrome – MSA + predominant autonomic dysfunction
-
SND – striato-nigral degeneration – older term, not much use - considered parkinsonism without response to levodopa without other features above
-
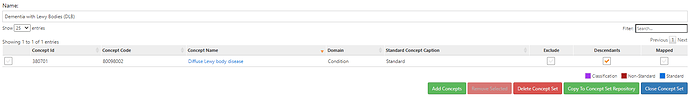
LBD/DLB – parkinsonism + early dementia with fluctuating mental status, hallucinations and sensitivity to side effects of dopaminergic medications
-
LBD – Lewy body dementia or DLB (dementia with Lewy bodies) are considered part of the spectrum of parkinsonism – this is controversial, though for phenotyping neurodegenerative parkinsonism, it is reasonable to include within the “neurodegenerative parkinsonism” group that:
Often “atypical parkinsonism” or “parkinsonism” are used by neurologists to mean neurodegenerative parkinsonism though the specificity of the terms itself is often in question.
- “parkinsonism” is often used in documentation (and in coding) to identify the syndrome without committing oneself to the diagnosis of PD (either due to lack of findings, progression, and sometimes due to lack of experience or confidence in making the diagnosis). And unfortunately many EHR systems map “parkinsonism, not specified” to G20 (ICD10: Parkinson’s disease, primary parkinsonism), reducing confidence in G20 being as specific to PD as it could be.
Here’s a nice simple overview (1 page!) provided by the Parkinson’s Foundation for parkinsonism and/vs Parkinson’s disease:
https://www.parkinson.org/pd-library/fact-sheets/parkinsonism-vs-parkinsons-disease
A basic overview of these conditions are in the same above PD reference as well:
Armstrong and Okun (2020 Diagnosis and Treatment of Parkinson Disease: A Review - PubMed )
Phenotype development
Here’s the practical “clinician” oriented summary of the clinical diagnosis considerations above:
This hierarchy is not matched perfectly to SNOMED/OMOP-CDM hierarchy which follows.
- Parkinsonism syndrome
**1.1. neurodegenerative parkinsonism (this is of interest #2) **
1.1.1. Parkinson’s disease (this is of interest #1)
1.1.2. non-PD neurodegenerative parkinsonisms (aka “atypical parkinsonism”)
1.1.2.1+ list of these: PSP, CBD, MSA (MSA-c, MSA-p, SDS), LBD, DLB, SND….
1.2. non-neurodegenerative parkinsonism (not of interest, but are often in the differential diagnosis under active consideration and if definitive and if the sole condition can be exclusionary); also known as “secondary parkinsonism”
1.2.1+. examples: secondary parkinsonism – drug-induced parkinsonism, vascular/multiinfarct parkinsonism, normal pressure hydrocephalus, postconcussive parkinsonism, postencephalitic parkinsonism, etc
Translating this to vocabulary considerations:
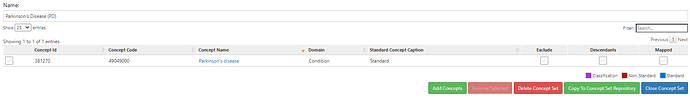
• Parkinson’s disease (specific)
• PD code alone: ICD10 code: G20
• OMOP standard: PD condition: 381270
• Parkinsonism codes (neurodegenerative parkinsonism)
• ICD10 codes: G20, G90.3, G23.1, G23.2, G31.83, G31.85, (G23.3)
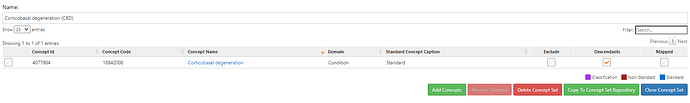
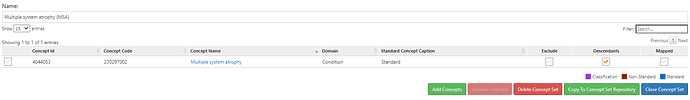
• OMOP standard concepts: PD, MSA, MSA-P, MSA-C, SDS, SND, parkinsonism w/orthostatic hypoTN, CBD, PSP, LBD
• Non-neurodegenerative parkinsonisms
• secondary parkinsonism: ICD10 codes: G21.*
• vascular parkinsonism:
• normal pressure hydrocephalus:
• drug-induced parkinsonism
And here’s the hierarchy of OMOP-CDM vocabulary codes mapped to ICD10 codes available
Literature on algorithms:
Review of many algorithms comes down to extraction of common elements with highly variable applications in several different combinations.
A good summary of 18 studies looking at algorithms for PD and parkinsonism (Harding et al 2019: Identifying Parkinson's disease and parkinsonism cases using routinely collected healthcare data: A systematic review - PubMed )
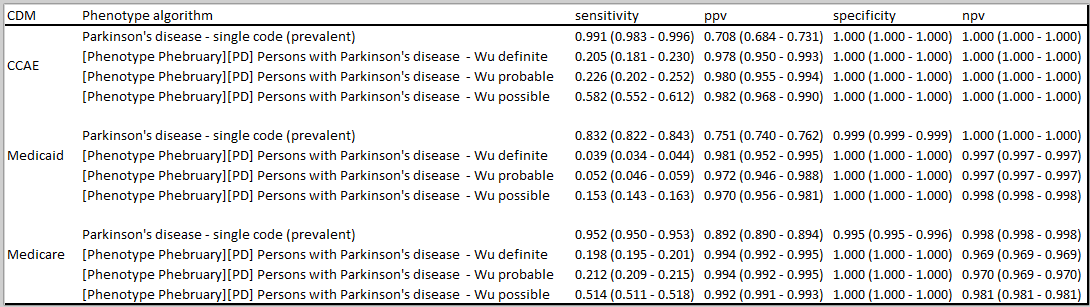
You can see a wide variety of PPV and sensitivity measures – and likely due to differing nature of cohorts, and as we would suspect, differing criteria for defining cases (PD or parkinsonism). Several of the broader ones also include secondary parkinsonism, mostly because there is a desire to capture broadly early cases of PD when it is often misdiagnosed or confused with secondary parkinsonism. Many used chart review as gold standard classification with some relying on neurology examination (this is another topic – how to standardize validation chart reviews across sites that our group is working on).
The CDC has created the National Neurologic Conditions Surveillance Survey (NNCSS) with PD as one of the first use cases (https://www.cdc.gov/surveillance/neurology/index.html). In discussions with CDC (not published or available), a comprehensive review of algorithms seeking a case definition for PD revealed more than 120+ algorithms published. This is an opportunity to leverage the OMOP-CDM community to start the process of comparing algorithms in a uniform scalable way will be an important contribution. Our group is helping advise CDC on this process.
The elements that are most often used for PD / parkinsonism algorithms are as follows:
• Diagnosis conditions that support dx:
o Specific PD diagnosis
o Broader parkinsonism diagnoses
• Diagnosis conditions that represent competing or alternative diagnoses that, when are confirmed would exclude PD/neurodegenerative parkinsonism, but may co-exist or be an incorrect diagnosis early in course
• Diagnosis position (primary diagnosis or not for that encounter)
• Encounters (ambulatory only or ambulatory/inpatient/ED inclusive)
• Specialty coding the diagnosis condition
o Neurologist coding diagnosis conveys more confidence in diagnosis code
o Movement Disorder neurology subspecialty (not in OMOP-CDM vocab) conveys even further confidence in diagnosis coding
• Medications that support dx
o Specific PD dopaminergic medications (levodopa ingredient and dopamine agonists)
o Other PD medications fairly specific to PD (amantadine, COMT-inhibitors, MAO-B inhibitors, a few others)
All of above feature a look-back time (typically 1-5 years).
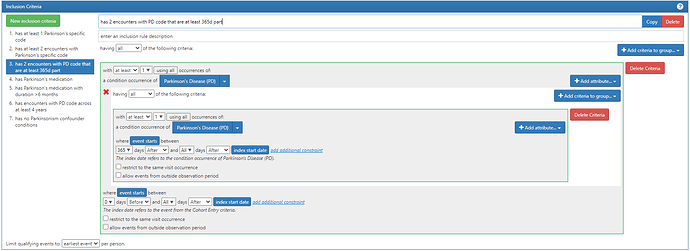
For diagnosis codes – typically # of coding events within lookback time as a threshold
For medications – either
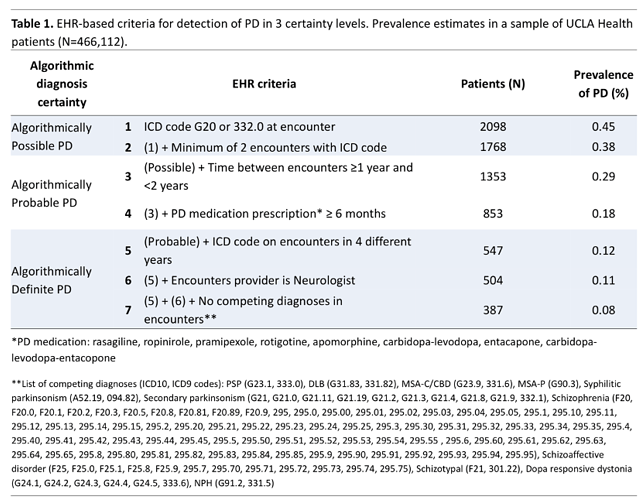
After manually reviewing all of criteria above in our local dataset, our group has presented an abstract proposing a tiered approach to assess effect of criteria: Folie et al Abstract presented at Movement Disorders Society 2021.
Recommended approach for OMOP Phenotyping exercise
Focus on a tiered approach for a systematic assessment toward specific PD while keeping a broad tier for parkinsonism (neurodegenerative).
Using above abstract as the framework, we note that we had focused on PD and not neurodegenerative parkinsonism, so it excluded specific neurodegenerative parkinsonism syndromes.
So there is need to add/create a separate set of tiers that do include a broader range of parkinsonism.
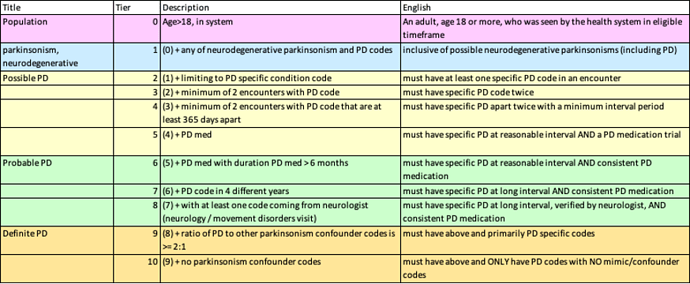
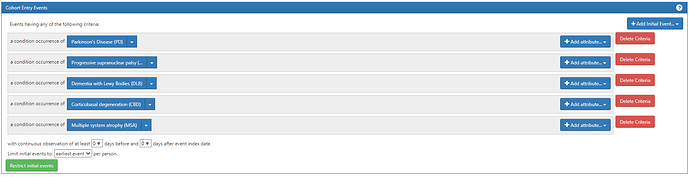
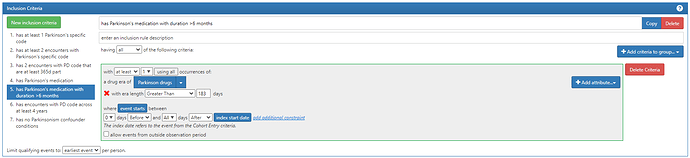
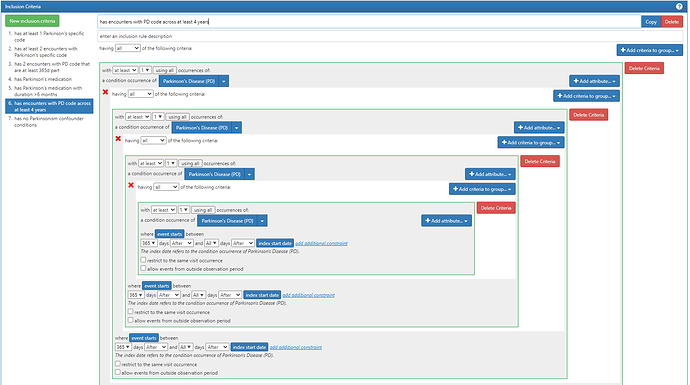
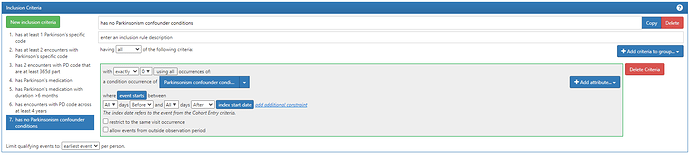
So we propose investigating the following tiers which we would recommend for the OMOP Phenotype February consideration:
algorithm cascade omop.xlsx (9.6 KB)
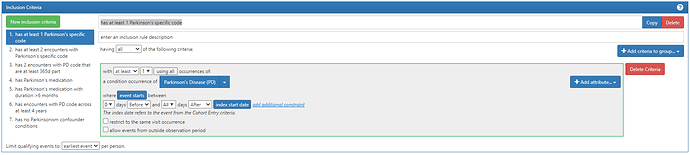
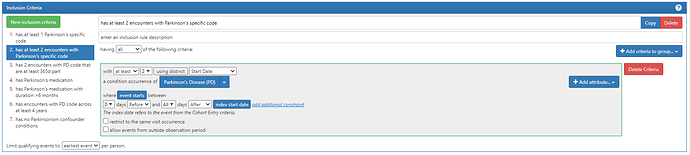
Inclusion diagnoses:
ICD10/9 code for PD: ‘G20’, ‘332.0’
ICD codes related to neurodegenerative parkinsonism (ICD10/9) includes ‘G23.1’, ‘333.0’, ‘G31.83’, ‘331.82’, ‘G23.9’, ‘331.6’, ‘G90.3’,‘G23.2’, ‘G23.3’,‘G31.85’
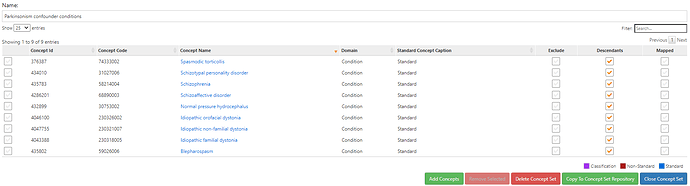
The ICD codes related to red flags (confounder codes) includes ‘F20’, ‘F20.0’, ‘F20.1’, ‘F20.2’, ‘F20.3’, ‘F20.5’, ‘F20.8’, ‘F20.81’, ‘F20.89’, ‘F20.9’, ‘295’, ‘295.0’, ‘295.00’, ‘295.01’, ‘295.02’, ‘295.03’, ‘295.04’, ‘295.05’, ‘295.1’, ‘295.10’, ‘295.11’, ‘295.12’, ‘295.13’, ‘295.14’, ‘295.15’, ‘295.2’, ‘295.20’, ‘295.21’, ‘295.22’, ‘295.23’, ‘295.24’, ‘295.25’, ‘295.3’, ‘295.30’, ‘295.31’, ‘295.32’, ‘295.33’, ‘295.34’, ‘295.35’, ‘295.4’, ‘295.40’, ‘295.41’,‘295.42’, ‘295.43’, ‘295.44’, ‘295.45’, ‘295.5’, ‘295.50’, ‘295.51’, ‘295.52’, ‘295.53’, ‘295.54’, ‘295.55’ , ‘295.6’, ‘295.60’, ‘295.61’, ‘295.62’, ‘295.63’, ‘295.64’, ‘295.65’, ‘295.8’, ‘295.80’, ‘295.81’, ‘295.82’, ‘295.83’, ‘295.84’, ‘295.85’, ‘295.9’, ‘295.90’, ‘295.91’, ‘295.92’, ‘295.93’, ‘295.94’, ‘295.95’, ‘F25’, ‘F25.0’, ‘F25.1’, ‘F25.8’, ‘F25.9’, ‘295.7’, ‘295.70’, ‘295.71’, ‘295.72’, ‘295.73’, ‘295.74’, ‘295.75’, ‘F21’, ‘301.22’, ‘G24.1’, ‘G24.2’, ‘G24.3’, ‘G24.4’, ‘G24.5’, ‘333.6’
And including as further confounder codes related to secondary parkinsonism includes ‘A52.19’, ‘094.82’, ‘G21’, ‘G21.0’, ‘G21.11’, ‘G21.19’, ‘G21.2’, ‘G21.3’, ‘G21.4’, ‘G21.8’, ‘G21.9’, ‘332.1’, ‘G91.2’, ‘331.5’
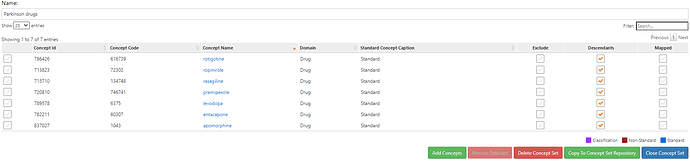
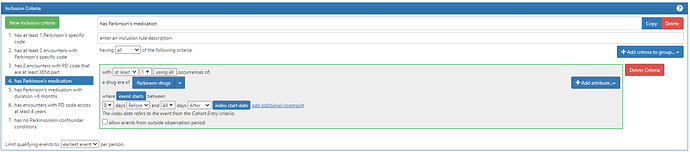
Proposed medication list for PD: any med with levodopa ingredient (typically forms of carbidopa/levodopa); pramipexole, ropinirole, rasagiline, rotigotine, apomorphine, entacapone
We likely will need to consider a separate cohort specification for developing a phenotype for specific non-PD neurodegenerative parkinsonisms, but we can start with above.
I would welcome input from this amazing community!
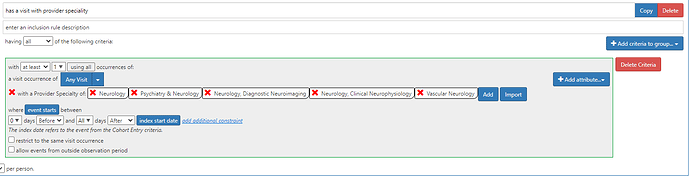

 ) which creates these sorts of challenges. Movement Disorders neurology is an commonly accepted specialty, but not an ACGME Fellowship supported one, so is basically not capturable within OMOP-CDM.
) which creates these sorts of challenges. Movement Disorders neurology is an commonly accepted specialty, but not an ACGME Fellowship supported one, so is basically not capturable within OMOP-CDM.