This post is part of a series showcasing my new R package Phea for electronic phenotyping.
How would you go about finding patients where the value of a measurement (e.g. body weight) has increased by at least X% in the past time period T?
Would a SQL join be the most efficient approach to write the query?
With Phea, this is how such a phenotype looks like:
phenotype <- calculate_formula(
components = list(
weight_current = weight_current,
weight_previous = weight_previous),
fml = list(
weight_change = '(weight_current_value_as_number - weight_previous_value_as_number) / weight_previous_value_as_number'))
The result phenotype is also a SQL query itself. You can keep just the rows where weight_change >= 0.5:
patients_50p <- phenotype |>
filter(weight_change >= 0.5) |>
select(pid) |>
distinct() |>
collect()
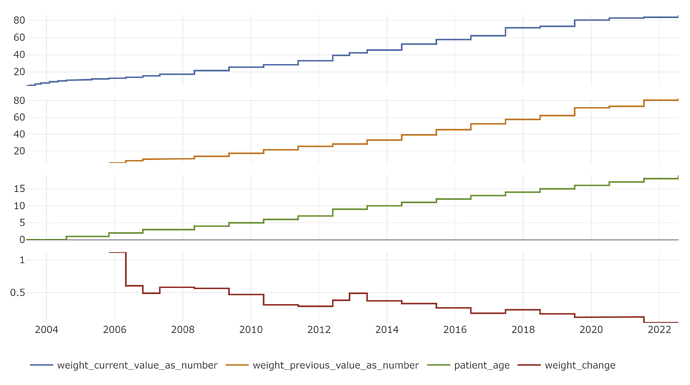
You can plot the phenotype for 1 patient that meets our criteria:
# Sample one patient at random
sample_patient <- sample(patients_50p, 1)
# Plot the phenotype for the sampled patient.
phea_plot(phen, pid = sample_patient)
And you can see the final SQL code of the query that calculates the phenotype here.
As it turns out, kids are good examples of patients whose weight increases by that much over 2-3 years ![]()
See full vignette at Calculate increase in body weight, or a primer on Phea here.
Kind regards!