Primary focus was building an ontology to support drug-related clinical decision support. This was for an EMR that included CUI as well as vocabulary-specific encoding.
For example, the NDDF has drug/disease indications and contraindications, drug/drug contraindications, drug/adverse event maps (e.g., monitor for fever or thrombocytopenia or elevated liver enzymes, etc.), routes of administration, etc. The ontology we built linked drug concepts to the symptom/sign/disease/lab concepts. None of these relationships were in the UMLS (MRREL, etc.) at the time we did the work.
We also did a fair amount of ingredient mapping for the ontology, so that we could easily identify all ACE inhibitors or all ARBs or all cephalosporins etc. for hypothesis testing prior to design of a pragmatic trial using EMR data.

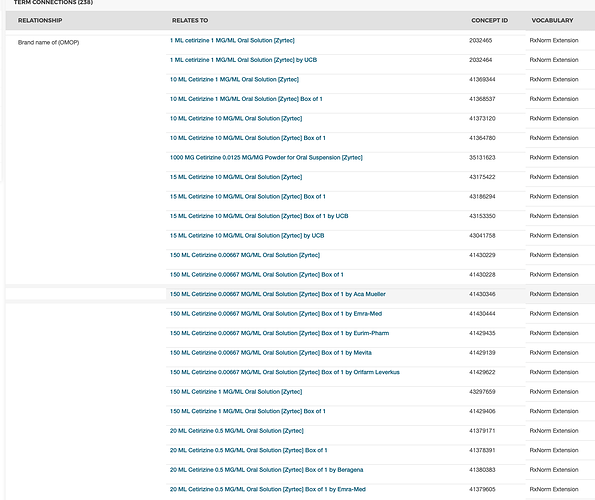
 I thought all RxNorm codes/concept_ids would map to a standard.
I thought all RxNorm codes/concept_ids would map to a standard.