- We preferred to map to Chinese version of Loinc at first. But there is no hierarchy information in Chinese version. Can anyone provide Chinese version of LOINC hierarchy concepts.
- For’ Lymphocyte absolute value’, we can’t find a exact concept in Loinc. Now we upward mapping to ‘Lymphocytes ’. Is this OK?
- For’ Hypersensitive C-reactive protein’, we can’t find a exact concept in Loinc. Now we upword mapping to ‘C-reactive protein’. Is this OK?
- For’ Percentage of B lymphocytes’, now we marked unable to map. Or can we map to ‘B lymphocytes’ or ’ lymphocytes percentage ’?
Is there any difference between the Chinese and international version? Why don’t you just use the hierarchy from the international one?
For the remaining questions: your LOINCs don’t look like LOINCs. For example, what is Lymphocytes? I know Lymphocytes [#/volume] in Blood by Automated count, Lymphocytes [#/volume] in Blood by Manual count and others but not just a generic “Lymphocytes”.
Regardless of that, you can go upward as you did or map to SNOMED measurements.
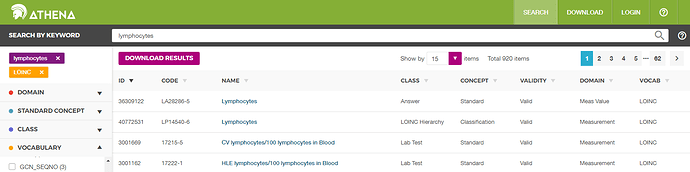
Until parts are imported into OMOP Vocabulary, you have to use search.loinc.org
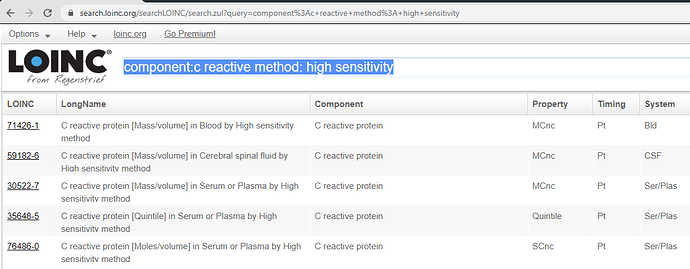
You can search by component. See blue highlight.
https://search.loinc.org/searchLOINC/search.zul?query=component%3Ac+reactive+method%3A+high+sensitivity
You don’t want the CSF code (System).
You can also use ranking here: ThemisConcepts/S20-ranking-LOINC-n-of-4-plus.csv at master · vojtechhuser/ThemisConcepts · GitHub
Some LOINC codes are “method-less”.
We preferred to map to Chinese version of Loinc at first. But there is no hierarchy information in Chinese version. Can anyone provide Chinese version of LOINC hierarchy concepts.
If the goal is to build OMOP CDM, it is more efficient to map the original data to international version of LOINC and then map to standard concept in OMOP. This can be achieved using the approach mentioned by @Vojtech_Huser. It would be cumbersome to first mapped to Chinese version of LOINC, then international
LOINC and then standard concept in OMOP. Don’t you agree?
Thanks for your help. I did the serach in http://athena.ohdsi.org/. And Lymphocytes [#/volume] in Blood by Automated count, Lymphocytes [#/volume] in Blood by Manual count has more information than ’ Lymphocyte absolute value’. We can’t do down mapping.
Thanks for your answer. I need to explain that ‘hypersensitive C-reactive protein,hs-CRP’ is a kind of C-reactive protein. ‘hypersensitive’ here is not a kind of method. By the way, what’s the CSF code. Hope your reply, thanks.
Yes, I agree. Now our problems about mapping is Chinese to English mapping needs more experience, time and expert people. Thanks a lot.
Most of the LOINC concepts are pretty specific and define not just an analyte, but method, sample, and property observed (concentration, mass, volume, etc.). Keep in mind that a time aspect of the measurement and its scale are also there, even though not visible in the concept_name. So you always need to consider the context of the lab test or map to generic SNOMED concepts.
Also, there is no way to map to the LOINC Answers (belongs to Meas value, not Measurement/Observation Domain) or LOINC Hierarchy classes (it’s Classification, not a Standard concept).
For ‘Lymphocyte absolute value’ use the SNOMED much more generic concept 40487382 Total lymphocyte count.
For ‘Percentage of B lymphocytes’ there is no option, but 4041971 B lymphocyte count.
I’m pretty sure ‘Hypersensitive C-reactive protein’ is the same protein detected by the highly-sensible method (hsCRP), so the options are:
46272908 Measurement of C-reactive protein using high sensitivity technique
2212762 C-reactive protein; high sensitivity (hsCRP)
Well, in this case depending on the figures you have in the source, look at 40762274 C reactive protein [Mass/volume] in Cerebral spinal fluid by High sensitivity method.