I’m trying to compile a list of sexually transmitted diseases using SNOMED-CT (I happen to be using OHDSI/OMOP concept tables in Databricks as my source for SNOMED-CT).
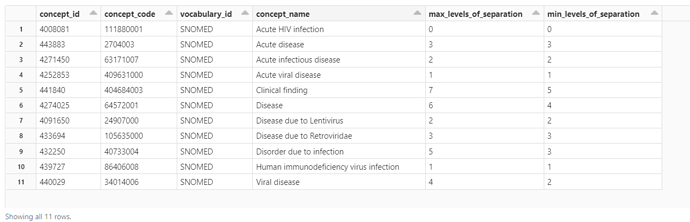
Querying the SNOMED-CT data from OHDSI Athena in OHDSI/OMOP using the query below, I get the results shown below. Notably missing from the ancestors is any indication that HIV is a sexually transmitted disease/infection.
Is there a way to use SNOMED-CT to create a somewhat comprehensive list of sexually transmitted diseases? Is there a better way to get to a list of codes (SNOMED or other, e.g. ICD) for sexually transmitted diseases/infections?
select distinct
parent.concept_id,
parent.concept_code,
parent.vocabulary_id,
parent.concept_name,
an.max_levels_of_separation,
an.min_levels_of_separation
from
concept con
join concept_ancestor an on 1=1
and an.descendant_concept_id = con.concept_id
join concept parent on 1=1
and parent.concept_id = an.ancestor_concept_id
where 1=1
and con.vocabulary_id = 'SNOMED'
and lower(con.concept_name) = 'acute hiv infection'
and con.domain_id = 'Condition'
order by parent.concept_name
;
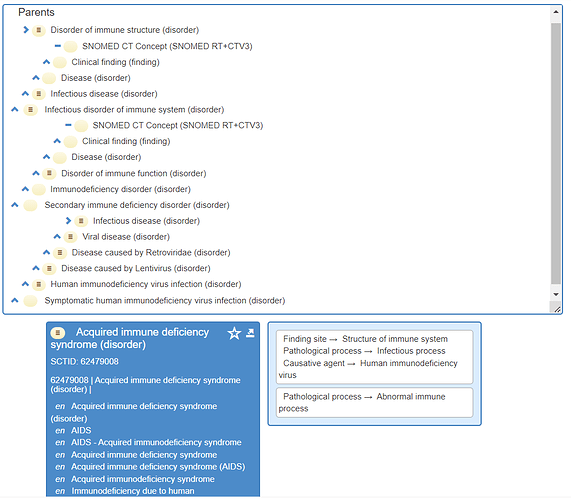
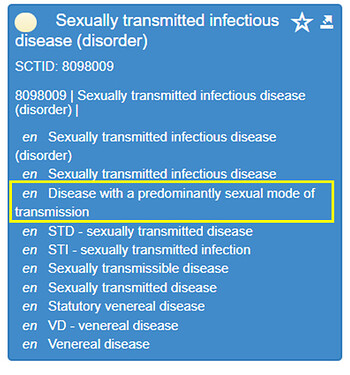
This finding seems to be confirmed using other SNOMED-CT browsers, for example: