Ah, so, @rkiefer, what you’ll need to do is create a new schema in your CDM database, and use the WebAPI endpiont: WebAPI/ddl/results?dialect=postgresql to get the DDL. You use this DDL to create the results schema tables. Note: there are a few table initialization scripts that will run forever if you do not have the proper indexes on your tables, please see this discussion for details:
We had run the results ddl previously and there are 82 tables in the schema. When I search the results ddl for postgresql, there is no create statement for the final_cohort table. What is the normal number of tables which should be created?
final_cohort table is a temp table in the generation script, which means it does not get created as a permanent table in the results schema.
The normal number of tables in the results schema are the tables that come out of the /ddl/results endpoint. It’s around 30 (not 81). Here is the list from my environment, note this may have a couple of extra tables compared to the version of the code that you are running (since the latest master has a few new results tables compared to version 2.6):
| TABLE_SCHEMA | TABLE_NAME |
|---|---|
| cknoll1 | ACHILLES_analysis |
| cknoll1 | ACHILLES_HEEL_results |
| cknoll1 | ACHILLES_results |
| cknoll1 | ACHILLES_results_derived |
| cknoll1 | ACHILLES_results_dist |
| cknoll1 | cc_results |
| cknoll1 | cohort |
| cknoll1 | cohort_censor_stats |
| cknoll1 | cohort_features |
| cknoll1 | cohort_features_analysis_ref |
| cknoll1 | cohort_features_dist |
| cknoll1 | cohort_features_ref |
| cknoll1 | cohort_inclusion |
| cknoll1 | cohort_inclusion_result |
| cknoll1 | cohort_inclusion_stats |
| cknoll1 | cohort_summary_stats |
| cknoll1 | concept_hierarchy |
| cknoll1 | feas_study_inclusion_stats |
| cknoll1 | feas_study_index_stats |
| cknoll1 | feas_study_result |
| cknoll1 | heracles_analysis |
| cknoll1 | HERACLES_HEEL_results |
| cknoll1 | heracles_periods |
| cknoll1 | heracles_results |
| cknoll1 | heracles_results_dist |
| cknoll1 | ir_analysis_dist |
| cknoll1 | ir_analysis_result |
| cknoll1 | ir_analysis_strata_stats |
| cknoll1 | ir_strata |
| cknoll1 | nc_results |
| cknoll1 | pathway_analysis_events |
| cknoll1 | pathway_analysis_stats |
In my env, my schema is cknoll1. You should have separate schemas for your CDM tables and your results tables.
Thanks @Chris_Knoll for your help. I worked with @rkiefer some more on this. In the logs of the Broadsea Docker image there was a nested exception which said this part of the query failed:
delete from ohdsi.cohort_inclusion_result where cohort_definition_id = 4 and mode_id = 0
This does not appear in the exported postgres SQL from ATLAS, when we were testing by hand.
The problem is that mode_id didn’t exist in the table. We cross-referenced against the WebAPI endpoint you noted (WebAPI/ddl/results?dialect=postgresql) and saw that mode_id was missing from a few tables. We manually fixed this, and queries are running now. Unsure how this happened, but will backtrack through our setup for ideas.
Right, there is a portion of the cohort generation in Atlas that generates the inclusion rule stats for reporting the inclusion rule impact reports (which is not necessary to do cohort generation, but is informative when looking at the report in Atlas).
Are you saying that the ddl/results script from the end point did not have a mode_id in the table definition? If so, I should make sure that the tables are properly scripted out, otherwise others will have this same problem.
The DDL scripts at the WebAPI endpoint did have the mode_id in the table definition. Sorry for the confusion. That gave us the right DDL so we could recreate the tables. We’re trying to trace back how the tables were initially provisioned as to why they were missing the column.
@lrasmussen @Chris_Knoll - Can the cohort generation fail due to lack of records matching my cohort definition (Say if my cohort entry criteria results in 0 records, can it fail?
For example, I created a concept set with ‘Warfarin’ and that has yielded results.
So when I used select * from codesets, I am able to find records but when I try the same after running “select * from qualified_events”, the result set is empty.
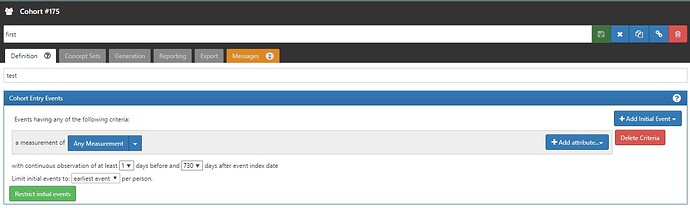
Can the cohort generation fail due to this? Moreover, I have set the cohort entry criteria as shown below
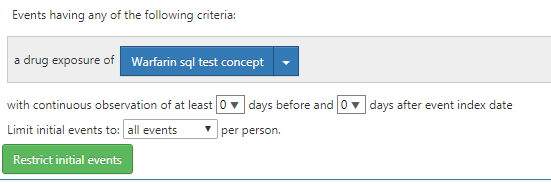
I tried with different settings for ‘iNITIAL EVENTS’ feature. But still am not able to generate the cohort successfully
We have data for Warfarin users. Can you people help us?
I tried running the queries seperately and they all are working fine. But certain queries yields empty table as results but I guess this is due to the criteria specified. Can cohort generation fail due to this?
Thanks
Selva
If you go to the SQL tab under the Export section (within cohort definitions), you can get the SQL that generates the cohort. You can execute it statement-by-statement to understand where you fail to return results.
Hello All,
I have set up Atlas at my end and one thing I observed that when I create new cohort and click generate, it says pending and remain stuck at that point.
I clicked Cancel, it doesn’t stop there, so I had to reboot the server.
Once rebooted, I click on Generate Cohort option for the same cohort which was showing pending earlier, runs within 7 seconds.
I am running it against SYNPUF 1k sample data.
Attaching the screenshot.
@Chris_Knoll @anthonysena @Ajit_Londhe (tagging you guys to get some help on this)
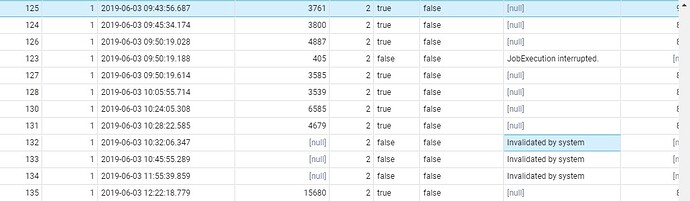
in webapi.cohort_generation_info table, I receive the following message in fail_message column “Invalidated by system”
I am using Postgresql.
TIA
‘Invalidated By System’ messages appears when you restart the service and there’s a job in a ‘running’ or ‘pending’ state. The jobs that were running between the restarts need to be reset (because the thread that is executing the analysis is killed, so on restart, all you can do is mark all running jobs ‘failed’ with ‘Invalidated by system’.
The original problem you were having is strange: you say that you click ‘generate’ but the job just sits in pending status? It’s possible you have another error happening between starting the job and the job starting the execution. Do you have anything in your logs? You mentioned that it did run successfully once. Can you get a simple cohort to generate or revert to the version that was working?
@Chris_Knoll
I deleted all the entries associated with cohort generation from webapi and results schema. Then I retried generating a simple cohort and facing the same issue with status “PENDING” from a long time. Attached the screenshot.
I am also attaching the screenshot for fail_message column.
I am on Atlas version 2.7.1 and WebAPI 2.7.1.
I will try to redo the cohort generation using previous version on WebAPI and Atlas.
Also, the cohort once generated doesn’t delete in one click, I had to reload the page again and again to make it work.
I don’ t see any logs which is associated with Atlas issues.
Kindly suggest.
Thank you
Hi,
I explained in the prior message: ‘Invalided by System’ mesasge happens when the WebAPI service is restarted and any job in ‘running’ state is reset with that failed message.
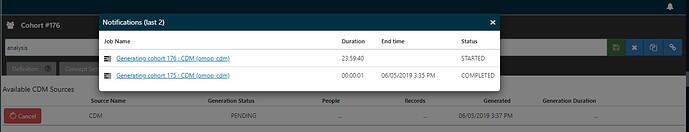
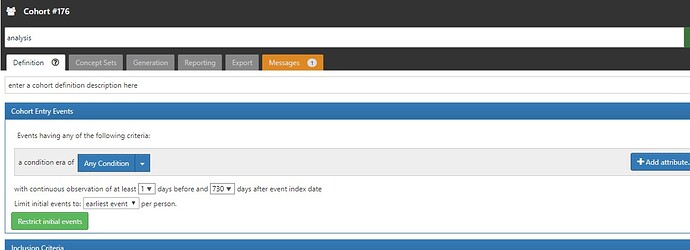
It looks like you did have a successful generation on cohort 175, so I’m not sure what is the issue with that cohort definition. Can you provide a screenshot of the cohort 176? Also, did you check your log files to look for an exception? Maybe there was an error in preparing the execution which causes the job to appear in ‘starting’ state but never actually starts executing due to some pre-launch error.
@Chris_Knoll
Hi Chris,
Here is what I did-
- Restarted tomcat service.
- Checked if WebAPI and Atlas are properly deployed and accessible via browser.
- Created Cohort # 175, clicked generate > failed
- Clicked generate again > complete
- Created cohort 176, clicked generate > PENDING > FAILED
- Clicked generate again > COMPLETED
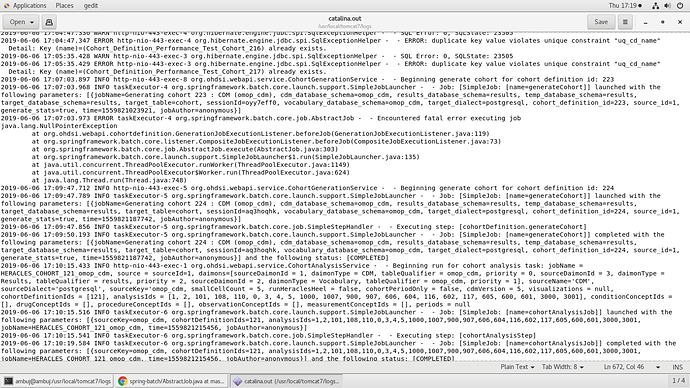
The logs that I can see is as below for the time when it failed:
Encountered one more scenario without touching WebAPI services:
Created cohort# 195> generate > Pending since past 15 mins
meanwhile the entry in DB table:
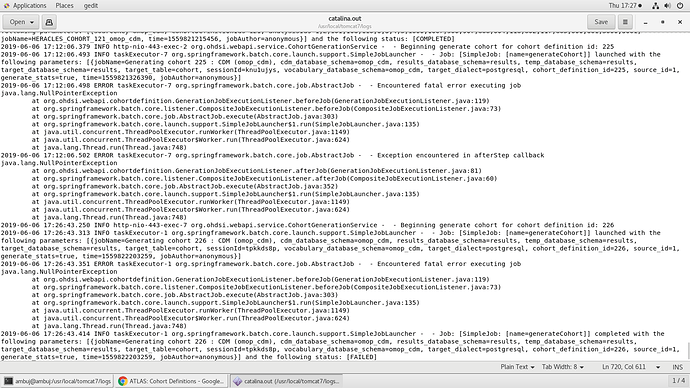
Additional Logs I found today, shows error when it gets stuck in PENDING state.
Below log is when it FAILED on Generate Cohort Click
Need your assistance on this.
Thank You
The issue is that the tables are created by the Flyway migrations. So, the cohort_inclusion_result table is created as follows:
Note the missing mode_id column. The same is true for the cohort_inclusion_stats and cohort_summary_stats tables.
The v2.5.0 release contains instructions on how to manually resolve the issue, but a Flyway migration was not included to automate this schema change. One possible solution is to provide a Flyway migration that brings the schema up to date, which is what I’ve done here.
For convenience, in Postgres, the solution is to run the following:
ALTER TABLE @resultsSchema.cohort_inclusion_result ADD mode_id int NOT NULL DEFAULT 0;
ALTER TABLE @resultsSchema.cohort_inclusion_stats ADD mode_id int NOT NULL DEFAULT 0;
ALTER TABLE @resultsSchema.cohort_summary_stats ADD mode_id int NOT NULL DEFAULT 0;
where @resultsSchema is the name of the schema containing the WebAPI tables.
Hi,
The tables created by flyway into the WebAPI database are no longer the approach to get the DDL for the results schema (hosted in the same database as your CDM schema).
The way to get the DDL for the results schema is to start your WebAPI instance, and open up the endpiont: /WebAPI/ddl/results?dialect={your dialect}. See SqlRender for your choice of dialects.
Since it may be difficult to understand which tables are new/altered from your current deployment, one way to figure it out is to deploy the latest results schema into a fresh/empty schema, and then use a DB schema compare utility to indicate which tables and columns are different between the new version and the current version. Then, you can plan your migrations based on your current situation.
Okay, good to know. Is that the long term plan as well? My use case might be atypical, but I am trying to automate the setup of a WebAPI v2.6.0 development environment, so I was hoping to avoid any manual steps. I ended up adding the migration so I could get my image to work (although I’ve only tested basic cohort generation). I’ll create a PR to show what I did.
EDIT: PR up here if anyone is interested:
Feel free to close it.
Yes, I think using the DDL is the long term. @JamesSWiggins uses it to automate deployments of WebAPI and Atlas onto the Amazon WebService platform. I’m pretty sure he uses the DDL directly out of the WebAPI endpoint.
We won’t be maintaining results schema via WebAPI in the future. In fact, we’d like to clean up those tables from the WebAPI schema, but we aren’t sure the impact that may cause, so we’re waiting for a 3.0 WebAPI release so that we can do some of that housekeeping.
-Chris
Hi,
Yes, please be encouraged to check out this automation in the GitHub repo below. Even if you aren’t deploying on AWS, the shell scripts contained within may be useful:
James
Hi All,
I am also facing a similar issue :
I am currently analyzing the features offered by the ATLAS tool. We are using a sample SynPuf data for this activity. We are using PostgreSQL based AWS RDS as the database.
Observation : The ATLAS tool created the cohorts and the reports were generated successfully. However, when we went to perform characterization and build cohort pathways, the process failed with the below error:
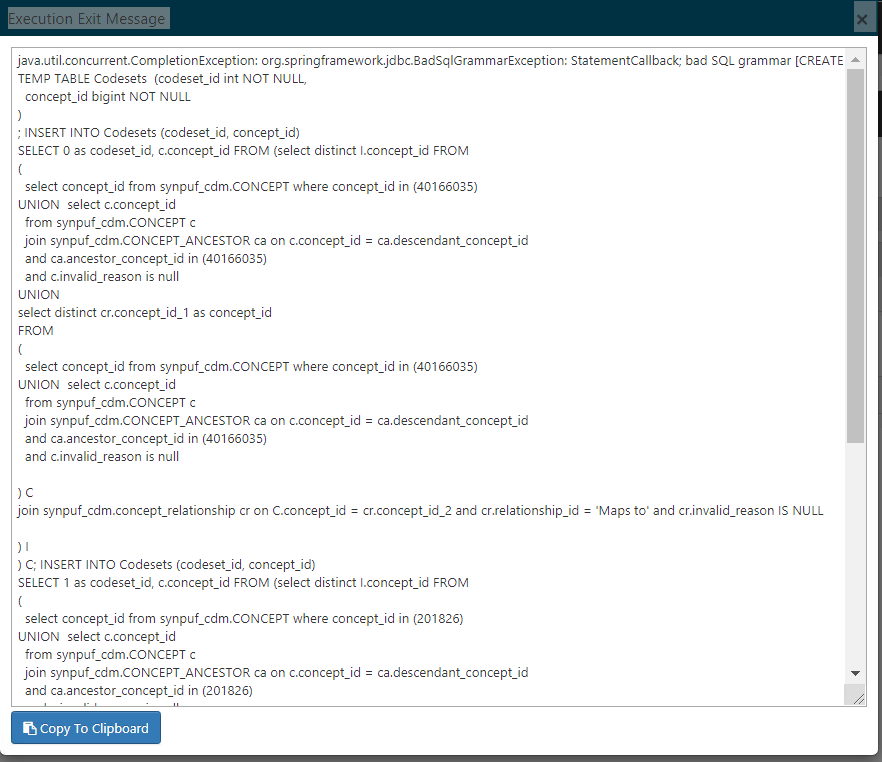
java.util.concurrent.CompletionException: org.springframework.jdbc.BadSqlGrammarException: StatementCallback; bad SQL grammar
Please refer to the image below :
We picked up the segment of queries mentioned in the error and executed it manually on our database and they executed without any issues.
We are looking for some help to resolve this. Please do let us know if you have any suggestions.
Thanks!
Thank you @psbrandt for posting this solution as I was coming up against this error. Also thanks to @Chris_Knoll and others on this thread as it has contained super useful information.
I have been using Broadsea (another thanks goes to @lee_evans) to run Atlas for the Synpuf data (CDM v5.3.1) contained in a Postgres database.
However (there’s always a but…) I run into issues when in Atlas on the Cohort Definitions page/Generation tab.
@psbrandt’s solution:
-- I use @resultsSchema = ohdsi as that seems to be what Broadsea uses as the results schema
ALTER TABLE ohdsi.cohort_inclusion_result ADD mode_id int NOT NULL DEFAULT 0;
ALTER TABLE ohdsi.cohort_inclusion_stats ADD mode_id int NOT NULL DEFAULT 0;
ALTER TABLE ohdsi.cohort_summary_stats ADD mode_id int NOT NULL DEFAULT 0;
removes the initial error when clicking Generate for the Cohort Definition, but now clicking Generate gives a new error:
# from the Generation Status FAILED hyperlink:
java.lang.RuntimeException: java.util.concurrent.ExecutionException: org.springframework.dao.DuplicateKeyException: StatementCallback; SQL [CREATE TEMP TABLE Codesets (codeset_id int NOT NULL,
concept_id bigint NOT NULL
)
; INSERT INTO Codesets (codeset_id, concept_id)
SELECT 0 as codeset_id, c.concept_id FROM (select distinct I.concept_id FROM
(
select concept_id from synpuf.CONCEPT where concept_id in (1312706,19087090,710062,715233,729855,740275,798874,711714,723013,43526424)
) I
) C; CREATE TEMP TABLE qualified_events
AS
WITH primary_events (event_id, person_id, start_date, end_date, op_start_date, op_end_date, visit_occurrence_id) AS (
-- Begin Primary Events
select P.ordinal as event_id, P.person_id, P.start_date, P.end_date, op_start_date, op_end_date, cast(P.visit_occurrence_id as bigint) as visit_occurrence_id
FROM
(
select E.person_id, E.start_date, E.end_date, row_number() OVER (PARTITION BY E.person_id ORDER BY E.start_date ASC) ordinal, OP... [truncated] ...t sr
CROSS JOIN (select count(*) as total_rules from ohdsi.cohort_inclusion where cohort_definition_id = 2) RuleTotal
where sr.mode_id = 1 and sr.cohort_definition_id = 2 and sr.inclusion_rule_mask = POWER(cast(2 as bigint),RuleTotal.total_rules)-1
) FC
; TRUNCATE TABLE best_events; DROP TABLE best_events; TRUNCATE TABLE cohort_rows; DROP TABLE cohort_rows; TRUNCATE TABLE final_cohort; DROP TABLE final_cohort; TRUNCATE TABLE inclusion_events; DROP TABLE inclusion_events; TRUNCATE TABLE qualified_events; DROP TABLE qualified_events; TRUNCATE TABLE included_events; DROP TABLE included_events; TRUNCATE TABLE Codesets; DROP TABLE Codesets]; ERROR: duplicate key value violates unique constraint "cohort_inclusion_stats_pkey"
Detail: Key (cohort_definition_id)=(2) already exists.; nested exception is org.postgresql.util.PSQLException: ERROR: duplicate key value violates unique constraint "cohort_inclusion_stats_pkey"
Detail: Key (cohort_definition_id)=(2) already exists.
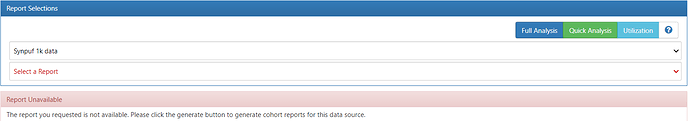
I can then get rid of this error by removing the cohort_inclusion_stats_pkey constraint for the cohort_inclusion_stats table in the ohdsi schema (which feels like a yucky thing to do!) in PGAdmin. There are subsequent errors for cohort_inclusion_* table constraints that I can delete also - then the Generation Status becomes “COMPLETE” in Atlas. However it seems to break the inclusion report that is meant to load (doesn’t load):
as well as giving a “The report you requested is not available. Please click the generate button to generate cohort reports for this data source” message on the Cohort Definitions/Reporting tab:
Is this a known issue/is there a solution/am I doing something wrong?
Also please let me know if you need other information about my setup (Win 10 Pro, Broadsea in Docker, Synpuf data (CDM v5.3.1) in a Postgres database).
Thanks for any help
Ty