Hello,
I’m trying to understand the intention of the source concept id field.
Is the source concept id designed to capture just the concept in the source data, or is it designed to capture both the clinical concept and terminology used natively within the source system to encode the data?
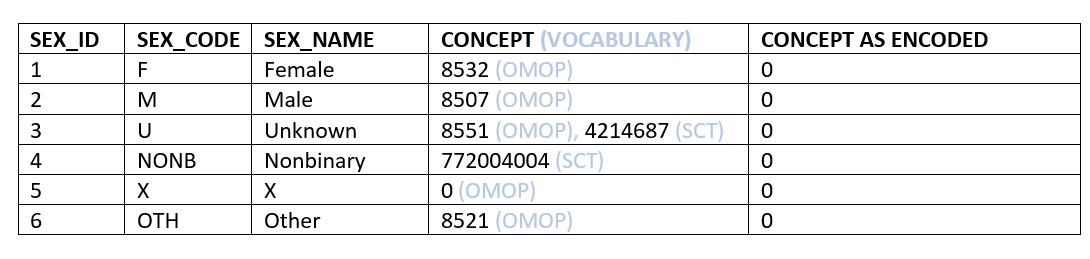
Being super concrete, if col 1-3 of the table below depicts the source data (encoded using organization specific codes), which codes are appropriate for the source concept id: col 4 - b/c just the source concept matters or col 5 - because we are making a claim about how the data is natively encoded (or not) in the source system:
Thanks in advance for your insight.